Promoter and operator play crucial roles in transcriptional regulation which is a vital process in all living organisms. The design of suitable promoter-operator transcriptional regulate system is the substantial work for constructing complex genetic networks. However, the existing of highly complex interactions between promoter and operator in natural biological systems made it hardly to fine-tune their functions independently.
The research group led by Prof. LOU Chunbo at the CAS Key Laboratory of Microbial Physiological and Metabolic Engineering, Institute of Microbiology, Chinese Academy of Sciences developed an insulate design principle adapted to prokaryotic transcriptional element. The principle realized the functional decoupling of promoters and operators in transcriptional control and enabled genetic circuit design with extraordinary precision and diversity.
In this research, by quantitative analysis, two sets of refined minimal promoters with high functional modularity were achieved, and the operator sequences containing unwanted transcriptional activity were excluded from the biological part library. A novel theoretical framework was developed for the parameterization of those parts and directing the manufacturing of transcriptional control system.
As a proof of those promoters and operators in the library has been well insulated and could be assembled predictably, 127 NOT gate circuits were constructed by randomly pairing the 53 promoters and 36 operators and tested in experiment. The data show that the measured output signals of combined circuits nicely match theoretical predicted signals.
Furthermore, these insulated transcriptional parts were utilized to build four-node transcriptional networks with incoherent feed-forward loop (IFFL) topology encoding a stripe-forming function. In silico screening of all 30,528 possible designs directly captured the one with the best stripe-forming performance without any trial-and-error work, as verified experimentally (up to 66-fold pulsing, mean errors <1.4-fold for all examined circuit designs). The results demonstrate the general applicability of the insulate design principle.
This insulation-based methodology simplifies the design-build-test-learn cycle of circuit engineering to a mix-and-match workflow, and enables computer-aided design of circuit functions in precision far exceeding those reported in previous studies. This research may inspire the modular design of other functional genetic parts; greatly enhance the ability of design and control more complex biological systems accordingly.
Their work was published on line in Nature Communications (http://www.nature.com/articles/s41467-017-00063-z) and is supported by the National Science Foundation of China and National Basic Research Program of China.
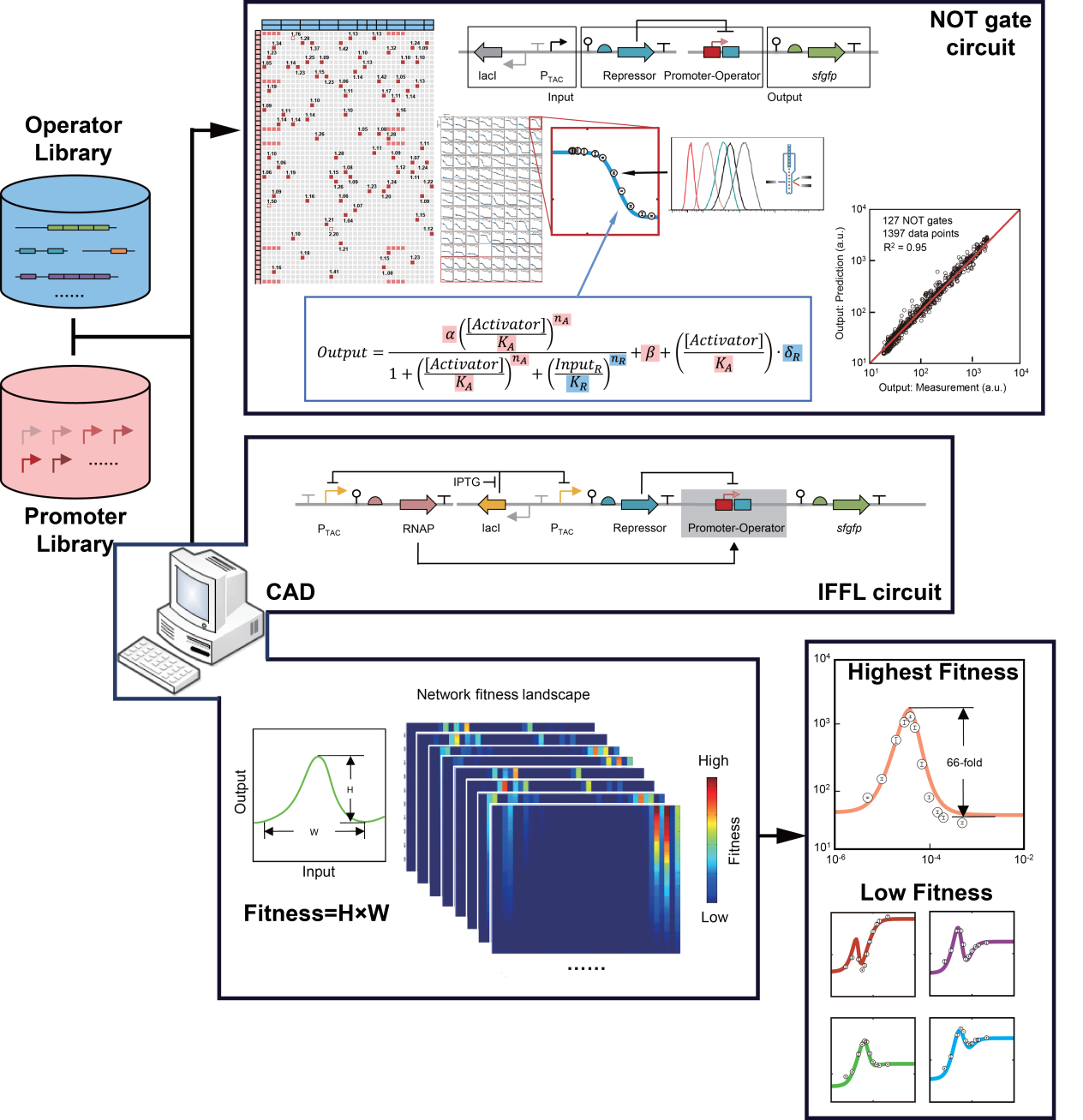
Overview of the precise design of genetic circuits using insulated promoters and operators. (Image by Prof. LOU Chunbo’s group)
CONTACT:
Dr. LOU Chunbo
Key Laboratory of Microbial Physiological and Metabolic Engineering, Institute of Microbiology, Chinese Academy of Sciences, Beijing, 100101, China
Email: louchunbo@im.ac.cn
