Genome editing has become increasingly important for biological and biotechnological research. Since the toxin-antitoxin (TA) systems in bacteria have evolved to facilitate the programmed cell death, techniques have been developed using only the toxin genes of the TA system as counter–selectable markers. However, these approaches are unstable and inefficient for genomic modifications of the model Gram-positive bacterium Bacillus subtilis due to the relative high rate of false positive mutants, which is a longstanding dilemma, has yet to be resolved.
The research groups of Profs. WEN Tingyi at the CAS Key Laboratory of Pathogenic Microbiology and Immunology, Institute of Microbiology, Chinese Academy of Sciences (IMCAS) and CHE Yongsheng at the State Key Laboratory of Toxicology & Medical Countermeasures, Beijing Institute of Pharmacology & Toxicology developed a new genome editing technology by reprogramming the bacterial TA systems as a toxin counter–selectable cassette regulated by an antitoxin switch (TCCRAS), which effectively introduced desired mutagenesis in the Gram-positive bacteria B. subtilis and Corynebacterium glutamicum. This work was published online in ACS Synthetic Biology (http://pubs.acs.org/doi/abs/10.1021/acssynbio.6b00287)
.
To mimic the TA behavior and design a toxin counter–selectable cassette regulated by an antitoxin switch (TCCRAS), the toxin and antitoxin components were reconstructed by the expression of toxins and antitoxin genes using the constitutive and inducible promoters, respectively. The RelBE complex, a member of the five characterized type II TA systems, was identified to specifically and efficiently regulate cell growth and death by the conditionally controlled antitoxin RelB switch, thereby serving as a novel counter selectable cassette to establish the TCCRAS system.
Using a single vector, this system has been employed to perform inframe deletion, functional knock-in, gene replacement, precise point mutation, large-scale insertion, and especially deletion of the fragments up to 194.9 kb in B. subtilis. In addition, the biosynthesis of lycopene was first achieved in B. subtilis using TCCRAS to integrate a 5.4-kb fusion cluster.
Additionally, the system was adapted for gene knockout and replacement, and large-scale deletion of the fragments up to 179.8 kb in C. glutamicum, with the mutation efficiencies increased by 0.8−1.0-fold compared to the conventional SacB method. The biosynthesis of cadaverine was achieved in the C. glutamicum, providing potential basis for the direct fermentation of cadaverine using C. glutamicum.
Taken together, a new genome editing technique TCCRAS was successfully established by constructing a toxin counter–selectable cassette regulated by an antitoxin switch, providing an effective and versatile genome-scale engineering technology for metabolic engineering and synthetic genomics research in a broad range of Gram-positive bacteria.
This method also served as an effective technology platform for constructing the artificial designed cell factories in the biosynthesis of new biocatalysts and important compounds, which would enhance the application value of microorganism in the field of medicine, agriculture and industry.
WU Jie, Ph.D. student and Dr. DENG Aihua from IMCAS are co-first authors of this paper, and Profs. WEN Tingyi and CHE Yongsheng are the corresponding authors.
This work was supported by the National Natural Science Foundation of China, National Hi-Tech Research & Development Program of China, Science and Technology Service Network Initiative, and the Key Deployment Projects of Chinese Academy of Sciences.
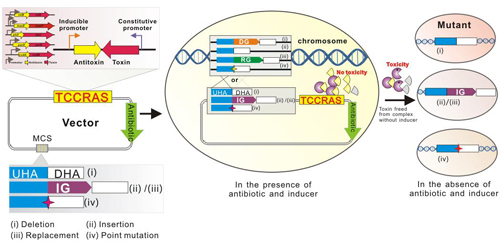
The schematic illustration of the genome editing using the TCCRAS system (Image by Prof. WEN Tingyi’s group)
Contact:
WEN Tingyi
CAS Key Laboratory of Pathogenic Microbiology and Immunology,Institute of Microbiology,Chinese Academy of Sciences,100101,Beijing, China
Phone: 86-10-64806119
E-mail: wenty@im.ac.cn
Key words:
genome editing, toxin-antitoxin (TA) system, Bacillus subtilis, Corynebacterium glutamicum
Abstract:
The bacterial toxin–antitoxin systems were reprogrammed as a toxin counter–selectable cassette regulated by an antitoxin switch (TCCRAS) for effective and versatile genome editing in the extensively studied and utilized Gram-positive bacteria, Bacillus subtilis and Corynebacterium glutamicum.
Reference:
Bacterial Genome Editing via a Designed Toxin−Antitoxin Cassette
http://pubs.acs.org/doi/abs/10.1021/acssynbio.6b00287
