2016.05.20, 4:00-5:00 PM, A-102
Spatially Interactive Biomolecular Networks Organized by DNA Architectures
Jinglin Fu 1,*
1 Department of Chemistry, Computational and Integrative Biology, Rutgers University-Camden, NJ08102, USA.
* Corresponding author: jinglin.fu@rutgers.edu
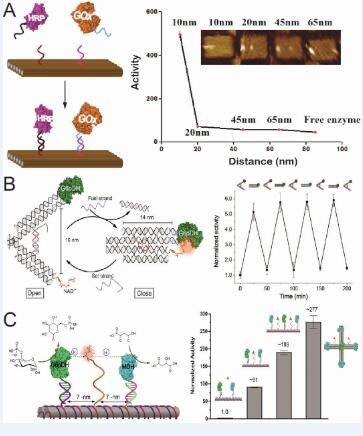
Living systems have evolved complex macromolecular nanostructure networks to mediate a range of cellular activities with high efficiency and specificity, including metabolic pathways, signaling transduction, gene expression and regulation. Many of these macromolecular systems are spatially organized with precisely controlled position and orientation of biomolecule components, which facilitate the functionality. If these systems can be mimicked and constructed artificially, they could be applied to the realization of smart nanobioreactors and devices that have utility in the production of high-value chemicals, the conversion of a variety of energy forms, and the development of new bio-diagnostics. These studies will also increase our fundamental knowledge of biomolecule selfassembly and cellular metabolism in general. Recently, DNA has emerged as promising scaffolds to organize macromolecule structures on the nanoscale with controlled geometries and nanomechanical capabilities.1 In this abstract, we present a DNA scaffold-directed strategy to assemble multi-enzyme systems with controlled spatial arrangements and interactions. In Fig.1A, GOx and HRP are organized with inter-enzyme distances ranging from 10 to 65 nm, confirmed by AFM imaging. The activity of assembled enzyme cascades decreased nonlinearly as increasing the distance between enzymes. The result directly demonstrates the ability of DNA nanostructures to arrange multiple proteins with control over their number, relative distance, and spatial pattern, mediating the substrate transport and activities. In Fig.1B, a tweezers-like DNA nanodevice is used to regulate the activity of an enzyme/cofactor pair. The actuation of enzymatic function was achieved by switching the tweezers between opened and closed states. Fig.1C demonstrates a biomimetic swinging arm capable of channeling substrates between enzymes. A two-enzyme cascade consisting of G6pDH and MDH was displayed on a DNA tile with a NADH arm positioned at halfway between two enzymes for transferring hydride intermediates. The activity of the G6pDH-MDH complex was enhanced by more than 90-fold with channeled NAD swinging arms.
References
(1) Zhao, Z., Fu, J.* et al. Nature Communications 2016, 7, 10619.
(2) Fu, J.; Liu, M.; Liu, Y. and Yan, H. Accounts of Chemical Research 2012, 8, 1215-1226.
(3) Fu, J.;* Yang, Y.; Johnson-Buck, A.; Liu, M.; Liu, Y.; Walter, N.; Woodbury, N. and Yan, H.* Nature Nanotechnology 2014, 9, 531–536.
(4) Liu, M.; Fu, J.;* Hejesen, C.; Yang, Y.; Woodbury, N.; Gothelf, K.; Liu, Y. and Yan, H.* Nature Communications 2013, 4, 2127.
(5) Fu, J.; Liu, M.; Liu, Y.; Woodbury, N. and Yan, H. Journal of American Chemical Society 2012, 134, 5516-5519.
Biography

Jinglin Fu graduated with Bachelor’s and Master’s degree in Chemistry from Zhejiang University. He received his Ph.D degree in Chemistry (2010) from Arizona State University with the thesis focused on peptides modulating enzyme functions. Then, he did a three-year postdoc with Prof. Hao Yan to use DNA nanostructures to direct the assembly of multi-enzyme systems. Currently, Dr. Fu is an assistant professor in Biochemistry at Rutgers University-Camden. Dr. Fu is also a joint member of Center for Computational and Integrative Biology at Rutgers. Dr. Fu is interested in the development of bio-inspired macromolecular complex that can function to control chemical catalysis, electron transfer and biomolecular interactions.
Host: Wenbin Du
