Nowadays, the fast and reliable in vitro detection of featured nucleic acid sequences is urgently demanded in health care, as well as food and environmental safety monitoring. The current in vitro nuclear acid detection method mainly relies on Polymerase chain reaction (PCR), to achieve detection. One needs specially designed primers for specific amplification or reporter systems, for instance, molecular beacon, to read internal sequence information of amplified fragments. Aforementioned techniques are faced with some limitations, such as a long development cycle, difficulty in exploit, need for costly equipment, poor signal output, as well as the difficulty in distinguishing nonspecific amplification products.
To solve these problems, Prof. LOU Chunbo’s group at the Institute of Microbiology, Chinese Academy of Sciences (IMCAS), together with the Peking iGEM team, developed a novel paired dCas9 reporter system. As a proof of concept, they successfully distinguished Mtb genome markers, thus established a universal pathogen detection system, providing new approaches for the development of in vitro nuclear acid detection.
At its core, researchers fused split halves of luciferase to a pair of dCas9 proteins to form a specific reporter system. Paired dCas9 reporter system is programmed by single guide RNAs complementary to the upstream and downstream segments of a target DNA sequence, respectively. When substrate DNA containing the two segment-sequences in proximity are detected and bound by a pair of dCas9, luminescence is generated from the catalytic activity of luciferase in its entirety.
In this detection system, sequence information (including complementary sequence and spacer sequence) and output signal amplification are obtained simultaneously. According to the programmability of CRISPR system, researchers mapped out a large pool of specific candidate sequences as the markers for Mtb H37Rv genome against the human oral metagenome. They then selected a portion of marker sequences among the large pool to execute high-throughput screenings, and succeeded detecting pathogen DNA, eventually demonstrated the possibility of developing an in vitro pathogen detection system.
This work has been published online in ACS Synthetic Biology. ZHANG Yihao, QIAN Long from Peking University and WEI Weijia from University of Chinese Academy of Sciences are co-first authors of this article. OUYANG Qi from Peking University and LOU Chunbo from IMCAS are co-corresponding authors. Dr. ZHAO Xuejin, Prof. LIU Cuihua from IMCAS provided technical assistance, other members of Peking iGEM team did lots of experimental work. LOU Chunbo’s lab is now optimizing and upgrading the Paired-dCas9 system to enrich its output signal pattern, to improve detection efficacy, as well as to simplify detection equipment and operating environment.
Hyperlink of this article: http://pubs.acs.org/doi/abs/10.1021/acssynbio.6b00215
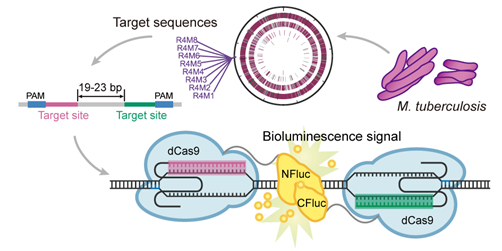
Fig. Paired Design of dCas9 as a Systematic Platform for the Detection of Featured Nucleic Acid Sequences in Pathogenic Strains (Image by LOU Chunbo)
Contact:
Dr. LOU Chunbo
CAS Key Laboratory of Microbial Physiological and Metabolic Engineering,Institute Of Microbiology,Chinese Academy of Sciences,100101,Beijing, China
E-mail: louchunbo@im.ac.cn