There is a consensual perspective that the interaction among microorganisms constitutes a driving force for fungi to produce numerous novel, complex, and diverse bioactive secondary metabolites (SMs), as abundance resources for drug discovery.
However, how do fungi communicate with microorganisms, and what are the molecular and biochemical mechanisms? Answers to these questions will foster a more detailed understanding of the complex ecological community on earth.
Dr. YIN Wenbing 's group screened out an endophytic fungus Epicoccum dendrobii which stimulated global alteration in the secondary metabolic profile of A. nidulans (Figure 1), during the exploration of interaction between the model fungus Aspergillus nidulans and other fungi.
Further research revealed that E. dendrobii was able to induce SM changes of at least fungi from four genera during cocultivation (Figure 1), indicating that the donor fungus E. dendrobii might contribute a general mechanism to trigger the recipient fungi which produce novel SMs by global regulations on secondary metabolism as the responses.
But how is this response achieved The researches demonstrated the response mechanism by chemical and genetic experiments during cocultivation of E. dendrobiiand A. nidulans. Transcriptome and metabolome analysis showed the widespread alteration of transcription and secondary metabolism in A. nidulans triggered by E. dendrobii, leading to the up-regulated expression of 15.4% and down-regulated of 19.0% of the genes in A. nidulans, as well as the up-regulation of 22 SMs which includes 8 novel polyketides (aspernidines) synthesized by the silent pkf BGC (Figure 1).
Unprecedentedly, gene deletion and replacement proved that the partial loss-of-function VeA1 protein is crucial for the SM alteration in fungal-fungal cocultivation.
Then, the velvet complex regulators VeA1, LaeA, and VelB but not VeA or VosA were proved to involve in the positive regulation of aspernidine derivatives during cocultivation. Using transcriptome analysis and gene deletions of potential transcription factors, the sole Zn(II)2Cys6 transcription factor SclB was confirmed to affect SM production in A. nidulans under cocultivation conditions.
In addition, the generalizability of regulatory network were confirmed in A.fumigatus and E. dendrobii cocultivation system by gene deletion and complementation.

Figure 1. The general cocultivation systems trigger by E. dendrobii (A and B); The widespread alteration of transcription and secondary metabolism in A. nidulans triggered by E. dendrobii(C and D) (Image by Dr. YIN’s group)
Cocultivation of E. dendrobii with A. nidulans carrying the veA1 allele led to widespread alteration of secondary metabolism, representing a new example for pathway activation. Partial loss-of-function of VeA (VeA1) is crucial for fungal chemical signal transduction triggered by cocultivation.
Therefore, cocultivation with E. dendrobii stimulated the VelB-VeA1-LaeA complex in A. nidulans, which subsequently induced the expression of SclB for eventually activating the silent pkf BGC and leading to the discovery of cryptic SMs (Figure 1).
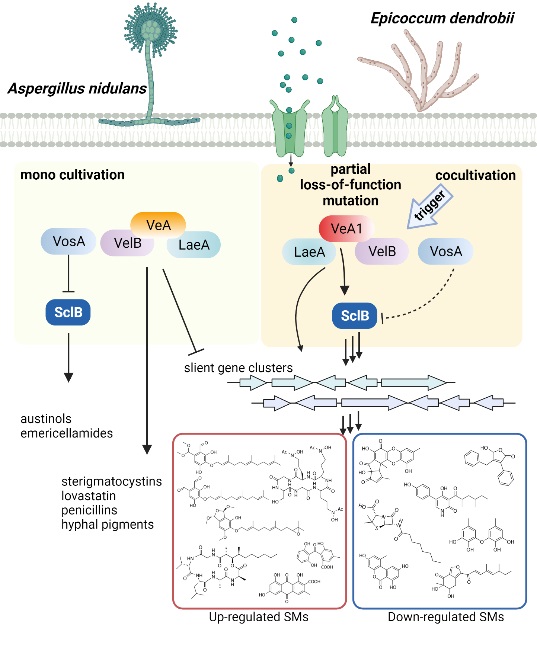
Figure 2. A model to show partial loss-of-function mutation of VeA-mediated secondary metabolism alterations during cocultivation of A. nidulans with E. dendrobii (Image by Dr. YIN’s group)
Full text links:
https://www.science.org/doi/10.1126/sciadv.abo6094
The paper entitled “Fungal-fungal cocultivation leads to widespread secondary metabolite alteration requiring the partial loss-of-function VeA1 protein” has been published online in Science Advances with Dr. WANG Gang and Dr. RAN Huomiao as joint first authors, Dr. YIN Wenbing as the corresponding author.
This work is jointly supported by the National Key Research and Development Program of China, the National Natural Science Foundation of China, the Key Research Program of Frontier Sciences, Chinese Academy of Sciences, the Biological Resources Program, Chinese Academy of Sciences, and the China Postdoctoral Science.
Contact:
Dr. YIN Wenbing
E-mail: yinwb@im.ac.cn
State Key Laboratory of Mycology, Institute of Microbiology, Chinese Academy of Sciences, Beijing 100101, PR China.